Approximately 1 in 2 Canadians will develop cancer in their lifetime. I am particularly interested in precision medicine for cancer patients. My research has focused on gene regulation in different cancer types and how this affects the behaviour of cancer cells. Ultimately, I aim to improve options and outcomes for individuals diagnosed with cancer.
What is gene regulation? Gene regulation controls the levels of a gene product in each cell in our bodies. We sometimes think of genes as “on” or “off”, but there are many different factors affecting the levels of these gene products that it’s more like a series of dimmer switches. Every step between DNA, messenger RNA, and protein can be modulated to affect the ultimate amount of gene products.
Why is gene regulation important for cancer? We usually think of cancer as a genetic disease that can be characterized by mutations (errors) in the DNA of cancer cells. Many of these mutations ultimately disrupt the regulation of genes that are important for normal cell behaviour. Characterizing the many layers of gene regulation in cancer cells can help scientists understand how to better treat patients with cancer.
To view my CV, you can click here: Krysta Mila Coyle .
My independent research program focuses on splicing as a mechanism of gene regulation in normal and malignant B-cells. I’m using isoform-based RNA-seq, bioinformatic pipelines, CRISPR-based prime/base editing, and other molecular & cell-based techniques to close the gap in the literature connecting specific alternative splicing events and global splicing aberrations to cancer biology, particularly in the study of lymphomas. I’ll be identifying biomarkers & testing novel therapeutic strategies. This work builds on the techniques and expertise developed throughout the selected projects below.
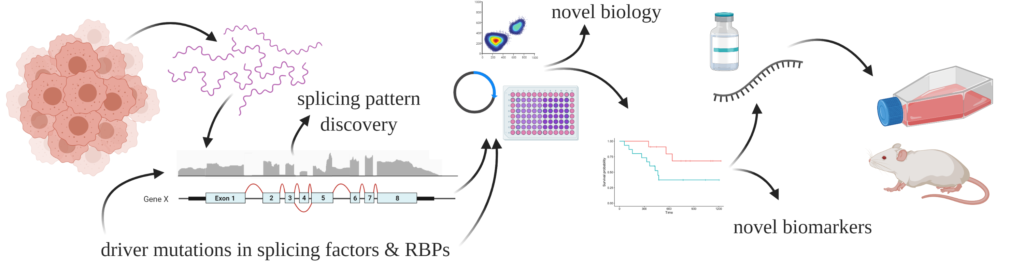
What are the impacts of non-coding mutations in mantle cell lymphoma?
While coding mutations in mature B-cell lymphomas have been described in detail, non-coding and silent mutations are often overlooked or ignored owing to a perceived lack of cellular function. We described a pattern of non-coding and silent mutations in the splicing factor HNRNPH1, which are associated with shorter patient survival. I used transcriptomics and molecular tools to demonstrate that these mutations result in altered splicing of HNRNPH1 and ultimately affect the abundance of the cognate protein. This led to a significant publication illustrating the combined importance of non-coding mutations, RNA binding proteins, and splicing in B cell lymphomagenesis (Pararajalingam and Coyle et al. Blood 2020).

Current work is focused on characterizing the splicing network associated with HNRNPH1 using clinical sequencing results as well as in vitro models of mantle cell lymphoma. I am also undertaking a pre-clinical investigation of an anti-sense oligonucleotide to correct the splicing of mutated HNRNPH1. Funding for this work has been provided by the American Society of Hematology and Lymphoma Canada.
How does genomic instability contribute to lymphomagenesis?
Mantle cell lymphoma (MCL) is a rare type of cancer that affects specific cells of the immune system, termed B cells. There is no cure for this type of cancer. MCL has many errors, or mutations, in elements that control the repair of damaged DNA. These mutations result in an relative inefficiency in repairing DNA damage in MCL. Consequently, MCL tumours accumulate many mutations and structural variants that can make the cancer more aggressive. We are studying a specific type of these errors that results when large sections of DNA shatter (chromothripsis or chromoplexy) and are reassembled in the wrong order. We hope that understanding how these catastrophes occur and are repaired will help us suggest new therapies for patients with mantle cell lymphoma.
Does canine B-cell lymphoma accurately reflect human disease?

Animal models of human cancers are an important tool for the development and preclinical evaluation of new treatments. Canine B-cell lymphoma (cBCL) is an appealing alternative to murine preclinical models due to its frequent, spontaneous incidence and its clinical and histological similarity to human B-cell non-Hodgkin lymphoma (NHL). The potential utility of cBCL as a veterinary model of human B-cell lymphomas would be bolstered by a more complete understanding of the genetic features found in cBCL. Using targeted sequencing of 86 canine patients, our study has revealed key differences in the mutational profiles of canine and human B-cell lymphomas and provides an impetus for enhanced genomic characterization of canine lymphomas as a model for human NHL, particularly in clinical trial settings. (Coyle and Hillman et al. Blood Adv. 2022.)
Does retinoic acid have use as a treatment for triple-negative breast cancer? (complete)
The first study investigated the role of a retinoid-inducible tumor suppressor, RARRES1, to determine whether it was a primary contributor to the response of breast tumors to retinoic acid. I used proteome and methylome analyses to demonstrate that regulation of RARRES1 varied by subtype of TNBC (Coyle et al. Oncotarget 2016). This finding was foundational to my work as it illustrated the importance of incorporating TNBC subtypes into further analyses. Furthermore, the majority of studies investigating retinoic acid in cancer have focused on retinoid metabolism pathways and canonical retinoid-responsive genes. I hypothesized that a more complex set of transcripts was involved in cellular responses to retinoic acid. To investigate this, I integrated methylome and transcriptome data to illustrate that the transcriptional response of breast cancer cell lines to all-trans retinoic acid was dependent on DNA methylation and primarily independent of the classical DNA retinoic acid response element (Coyle et al. Sci Rep 2017). The culmination of this work was a detailed in vivo study with cell line and patient-derived xenografts, integrating transcriptome and methylome data to evaluate the efficacy of retinoic acid as a potential therapeutic agent (Coyle et al. Cancers 2018).